Arrhythmias remain hard to predict
Cardiac arrhythmias are among the most dangerous manifestations of heart disease. They arise when the finely tuned electrical activity that coordinates the heartbeat becomes disrupted, sometimes leading to life-threatening events. Despite major advances in cardiac imaging and computational modelling, predicting when and why arrhythmias occur remains an unresolved challenge.
The ERC Advanced Grant project HeartCORE addresses this gap with an ambitious goal: to integrate high-resolution functional and structural data across the entire three-dimensional heart and translate this integration into predictive computational models. By unifying optics, cardiac electrophysiology, and computational modelling, HeartCORE seeks to reshape our understanding of arrhythmogenesis and open new perspectives for predictive cardiology.
Heart disease: when structure reshapes electrical behaviour
Cardiac remodelling is a hallmark of many heart diseases, both genetic and acquired. Excessive collagen deposition, changes in cellular orientation, and the reorganisation of cardiac non-myocytes can profoundly alter the pathways through which electrical impulses propagate. These structural alterations are strongly associated with conduction disturbances and arrhythmias. Yet, the precise mechanisms through which they unsettle electrical activity remain elusive.
Existing computational models often struggle to reproduce experimentally observed electrical behaviour in remodelled hearts. They are usually built on low-resolution functional measurements or simplified anatomical representations, limiting their ability to capture the multi-scale nature of cardiac tissue. This raises a fundamental question with both scientific and clinical relevance: what level of structural detail is necessary to predict electrical dysfunction in the heart?
The HeartCORE concept: integrating function and anatomy
The HeartCORE project is built on the premise that only by integrating function and structure at comparable resolutions can we uncover the true dynamics linking cardiac remodelling to arrhythmias. Through an interdisciplinary workflow combining advanced optical imaging techniques with organ-scale structural reconstruction and data-driven computational modelling, HeartCORE can truly uncover these dynamics. In this way, HeartCORE uniquely connects 3 domains traditionally studied separately: electrophysiology, tissue architecture, and computational modelling.
First, the electrical activity of entire hearts is characterised using innovative optical mapping techniques that capture conduction patterns and arrhythmogenic behaviour with high spatial and temporal resolution in 3D. Unlike conventional approaches that observe the heart from a single viewpoint, HeartCORE employs panoramic and multi-depth imaging strategies to obtain a volumetric representation of cardiac electrical activity.
Next, the same hearts are transformed into optically transparent organs and imaged at mesoscopic resolution. This step enables the reconstruction of the 3D organisation of cardiac tissue, including muscle fibres and collagen networks, across the entire organ. Selected high-resolution histological and immunohistochemical data are integrated into these reconstructions to provide detailed information about cellular composition and tissue organisation.
Finally, functional and structural datasets are merged into computational models capable of simulating cardiac electrical activity. These ‘virtual hearts’ allow researchers to test whether observed electrical phenomena can be explained by structural features alone or whether additional electrophysiological mechanisms must be invoked.
By combining these elements into a single framework, HeartCORE establishes a new paradigm for studying the interplay between structure and function in the heart.
Optical challenges when imaging entire hearts
A central technological innovation of HeartCORE is the ability to link functional measurements obtained from living hearts with structural information derived from the same organs. This requires overcoming significant technical challenges, including the need for high-resolution three-dimensional imaging across large tissue volumes and the integration of heterogeneous datasets.
To address these challenges, the project employs advanced optical platforms capable of capturing electrical activity across the entire heart surface and through its depth. By exploiting different excitation wavelengths, researchers can distinguish signals originating from superficial and deeper tissue layers, providing unprecedented insight into the transmural dynamics of cardiac electrophysiology.
Following functional characterisation, hearts undergo tissue-clearing procedures that render them transparent while preserving their structural integrity. Combined with light-sheet microscopy, these techniques enable rapid imaging of whole organs at micrometric resolution. The resulting datasets reveal the intricate architecture of cardiac tissue, including fibre orientation, fibrosis patterns and the spatial distribution of different cell types.
This dual perspective—functional and structural—forms the foundation for the subsequent modelling phase, in which experimental observations are translated into predictive simulations.
Building a virtual heart
The integration of morpho-functional data culminates in the creation of computational models that simulate electrical propagation in realistic cardiac geometries. These models incorporate information about tissue structure, cellular orientation and electrophysiological properties, enabling the reproduction of experimentally observed conduction patterns.
A key innovation of HeartCORE is the inclusion of optical signal modelling alongside electrophysiological simulations. Because signals derived from optical mapping reflect combined activity across the ventricular wall, the project incorporates models of light propagation to bridge the gap between simulated electrical activity and experimentally measured optical signals. This approach enhances the accuracy of model calibration and validation.
The resulting virtual hearts serve as experimental in silico platforms. By systematically varying structural and electrophysiological parameters, researchers can explore the conditions under which arrhythmias emerge and identify the mechanisms responsible for conduction disturbances. Crucially, this framework allows the evaluation of whether structural remodelling alone is sufficient to explain arrhythmogenic behaviour or whether additional factors, such as changes in cellular electrophysiology or interactions between different cell types, play a decisive role.
Two clinically relevant scenarios
To demonstrate its translational potential, HeartCORE focuses on 2 clinically significant forms of cardiac remodelling.
Hypertrophic cardiomyopathy
In hypertrophic cardiomyopathy, genetic mutations lead to thickening of the heart muscle, cellular disarray and diffuse interstitial fibrosis. These structural changes are associated with an increased risk of arrhythmias and sudden cardiac death, yet the underlying mechanisms remain incompletely understood.
HeartCORE investigates this condition using a transgenic rabbit model that closely mimics human cardiac electrophysiology. By combining optical mapping with organ-scale structural reconstruction, the project examines how interstitial fibrosis and cellular disorganisation influence conduction patterns and arrhythmogenicity. Computational models are then used to test whether these structural features can account for observed electrical disturbances or whether additional electrophysiological heterogeneity is required.
Myocardial infarction
Myocardial infarction represents a contrasting scenario in which localised regions of compact fibrosis form scar tissue within the heart. Traditionally considered electrically inert, scars are now known to participate in complex conduction phenomena, including trans-scar propagation and altered conduction velocity at border zones.
HeartCORE explores these phenomena using rabbit models of infarction generated through different surgical interventions. By integrating functional and structural data, the project analyses how scar architecture and cellular composition influence electrical propagation. Detailed computational models incorporating cardiomyocytes, fibroblasts and other cell types are used to investigate the role of cardiac non-myocyte interactions in shaping electrophysiological behaviour.
Together, these 2 cases provide a comprehensive view of how distinct forms of remodelling—diffuse and localised—affect cardiac electrical dynamics.
Beyond current predictive models
One of the most significant implications of HeartCORE lies in its potential to challenge existing predictive paradigms in cardiac electrophysiology. Many contemporary approaches, including digital twin strategies, rely heavily on structural information to infer electrical behaviour. While these methods have shown promise, HeartCORE’s preliminary findings suggest that structural data alone may be insufficient to capture the full complexity of arrhythmogenesis (Giardini et al., 2025; Nature Cardiovascular Research, 2025).
By integrating high-resolution morpho-functional data with advanced modelling techniques, HeartCORE provides a framework for assessing the relative contributions of structure and electrophysiology to arrhythmic risk. This approach not only refines our understanding of cardiac remodelling but also offers a pathway towards more reliable predictive tools.
Moreover, the project highlights the importance of interdisciplinary collaboration in tackling complex biomedical problems. By bridging optics, physiology, mathematical modelling, and computational science, HeartCORE exemplifies how technological innovation can drive conceptual advances in medicine.
A new framework for understanding arrhythmogenesis
HeartCORE represents a bold step towards a unified understanding of cardiac structure and function. By correlating electrical activity with three-dimensional tissue architecture at unprecedented resolution, the project moves beyond descriptive correlations and towards mechanistic explanations of arrhythmogenesis.
The data and models generated within HeartCORE are expected to constitute a valuable resource for the scientific community, enabling the testing of hypotheses and the development of new predictive strategies. In the long term, this integrative approach may contribute to more personalised risk assessment and targeted interventions for patients with cardiac disease.
In redefining how we study the relationship between structure and electrical function in the heart, HeartCORE offers not only new insights into arrhythmias but also a blueprint for future research at the intersection of biology, physics, and computation.
References
Giardini, F. et al. (2025) ‘Correlative imaging integrates electrophysiology with three-dimensional murine heart reconstruction to reveal electrical coupling between cell types’, Nature Cardiovascular Research, 4, pp. 14661486. Available at: https://doi.org/10.1038/s44161-025-00728-9.
Nature Cardiovascular Research (2025) ‘Myocardial fibrosis promotes arrhythmias at fast pacing rates’, Nature Cardiovascular Research, 4, pp. 1462–1463. Available at: https://doi.org/10.1038/s44161-025-00738-7.
PROJECT SUMMARY
HeartCORE is an ERC Advanced Grant project that aims to improve the prediction of cardiac arrhythmias by integrating high-resolution functional and structural data at the whole-heart level. By combining advanced optical mapping, three-dimensional tissue reconstruction, and computational modelling, HeartCORE establishes a unified framework to investigate how cardiac remodelling shapes electrical dysfunction in clinically relevant disease scenarios.
PROJECT PARTNERS
HeartCORE is led at the Institute of Clinical Physiology of the National Research Council (CNR, Italy) and is supported by an interdisciplinary network of academic collaborators. Key partners include Elisabetta Cerbai (University of Florence), Francesco Regazzoni (Politecnico di Milano) and Godfrey Smith (University of Glasgow), contributing complementary expertise in cardiac electrophysiology, computational modelling, and translational cardiovascular research.
PROJECT LEAD PROFILE
Leonardo Sacconi is Research Director at the Institute of Clinical Physiology of the National Research Council (CNR, Italy). His research focuses on the development of advanced optical methods to study cardiac electrophysiology from the cellular to the whole-heart scale. He leads the OptoCARD research group, which combines biophotonics, cardiac physiology, and computational approaches.
PROJECT CONTACTS
Dr Leonardo Sacconi
Research Director – OptoCARD Lab Institute of Clinical Physiology (IFC) – CNR; Viale Gaetano Pieraccini, 6; 50139 Florence (Italy)
Email: leonardo.sacconi@cnr.it
Web: optocard.it
LinkedIn: /in/leonardo-sacconi-b302ab349/
FUNDING
This project has received funding from the European Research Council (ERC) under the European Union’s research and innovation programme (Grant agreement No. 101198778).
Views and opinions expressed are, however, those of the author(s) only and do not necessarily reflect those of the European Union or the ERC. Neither the European Union nor the granting authority can be held responsible for them.
Figure legends
Figure 1: Computational modelling represents complex systems through discretisation, where a continuous structure is divided into a mesh of elements. This approach enables numerical simulations that approximate physical phenomena and provide insight into system behaviour.
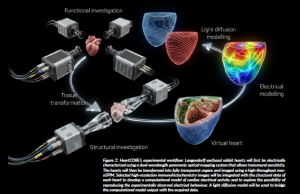
Figure 2: HeartCORE’s experimental workflow: Langendorff-perfused rabbit hearts will first be electrically characterised using a dual-wavelength panoramic optical mapping system that allows transmural sensitivity. The hearts will then be transformed into fully transparent organs and imaged using a high-throughput mesoSPIM. Selected high-resolution immunohistochemistry images will be integrated with the structural data of each heart to develop a computational model of cardiac electrical activity and to explore the possibility of reproducing the experimentally observed electrical behaviour. A light diffusion model will be used to bridge the computational model output with the acquired data.