Claudia Bartoli and Fabienne Vailleau, Université de Toulouse, INRAE, CNRS, LIPME,
Castanet-Tolosan, France
Coevolution between plants and their associated microbial communities is a central topic in evolutionary biology. Understanding how plant genomes evolve in interaction with their microbiota is essential to decipher the processes shaping plant adaptation. The HOLOE2Plant project addresses this challenge by investigating the coevolution between Brassica rapa and its associated microbiome, thereby contributing to the development of the holobiont concept, in which the plant and its microbiota function as a single evolutionary unit.
Beyond its fundamental relevance, this research has strong implications for agriculture. By studying both fungal and bacterial communities associated with natural B. rapa populations, we identified shared plant genetic bases involved in interactions across microbial kingdoms. These findings were recently reported in Plant Genetic Bases Associated With Microbiota Descriptors Shed Light Into a Novel Holobiont Generalist Genes Theory (Maillet et al., 2025). In this study, combining microbiome profiling, whole-genome sequencing and Genome–Environment Association (GEA) analyses across 26 natural B. rapa populations, we identified a set of 13 plant genes associated with both fungal and bacterial microbiota descriptors, which we defined as holobiont generalist genes (HGGs). Most of the identified HGGs showed traces of selection. These results provide strong empirical support for the idea that plants have evolved shared genetic mechanisms to regulate multi-kingdom microbial interactions, a key step forward in validating the holobiont framework in natural systems.
Studying natural populations and exploring their vast diversity
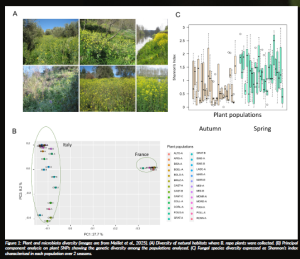
To investigate long-term plant–microbe coevolution, we focused on wild B. rapa populations in southern Italy and northern France. These populations were sampled across diverse habitats (Figure 1A), and their associated microbes were characterised using high-throughput sequencing technologies. In parallel, plant genetic diversity was assessed through whole-genome sequencing and bioinformatic analyses. We found that Italian populations exhibit higher genetic diversity than French populations (Figure 1B), likely reflecting the Mediterranean region as a centre of origin for Brassica species. Microbial communities were also highly diverse and strongly structured according to plant populations (Figure 1C), both in composition and species richness.
These results reinforce the idea that host plants actively shape their microbiota. Importantly, natural plant populations—unlike selected crop varieties—have coevolved with their microbial partners over long evolutionary timescales, making them powerful systems to uncover adaptive plant–microbe interactions. This is why working on wild plant species can help scientists better understand how hosts positively select for microbiomes that help them adapt to fluctuating environments and epidemics.
Linking microbial communities to plant genetic loci
Using a GEA approach, we correlated plant genetic variation to microbiota composition in root and rhizosphere compartments. This approach allows the identification of plant loci associated with environmental variables, including microbial descriptors such as species richness, composition or the presence/ absence of specific microbial species. Our analyses revealed that several plant loci are simultaneously associated with fungal and bacterial communities. The identification of HGGs demonstrates that plants may rely on integrated genetic pathways to interact with multiple microbial kingdoms, rather than independent mechanisms. Importantly, several of these loci show signatures of natural selection, highlighting their adaptive significance. Among them, we identified genes associated with microbial genera such as Bacillus and Trichoderma (Figure 2), well known for their plant growth-promoting and biocontrol properties. In addition, plant loci associated to these beneficial microbes show orthologues in other plant species such as Arabidopsis thaliana, making them excellent candidates for translational science. These first results from the HOLOE2Plant project suggest that natural plant populations have evolved the ability to precisely recruit beneficial microbes that enhance resistance to pathogens and, probably, to environmental stresses.
Learning from nature to reduce pesticide use
Plant genetic and microbial diversity are key drivers of plant resilience. Natural plant populations harbour a wide range of genetic variants enabling flexible interactions with diverse microbial partners under several environmental constraints. As a consequence, they are generally more resilient to diseases than modern crops, which often lack genetic diversity and are grown in poor microbiota soils. Leveraging this diversity opens new opportunities for sustainable agriculture and pesticide reduction. Within HOLOE2Plant, we established a large collection of bacterial and fungal isolates from diverse B. rapa populations and identified key strains capable of protecting plants against fungal diseases. We are now using experimental evolution approaches to understand how beneficial microbes can enhance plant defences over multiple generations, moving towards the design of stable and adaptive plant-microbiome systems. With large-scale greenhouse experimentation, we demonstrated that microbial communities from natural B. rapa populations can effectively protect the host against diseases.
What comes next: towards microbial-based solutions and precision agriculture
The next phase of our research aims to bridge evolutionary biology with applied agricultural innovation. Ongoing experimental evolution studies using fast-cycling B. rapa and ad hoc-designed microbial communities will allow us to: 1) track coevolutionary dynamics between plant genes (including HGGs) and microbial communities; and 2) identify how beneficial interactions become stable across generations.
In the longer term, these results will directly contribute to the development of next-generation agricultural systems combining microbial-based solutions inspired by natural communities and plant breeding for microbiome recruitment: selecting crop genotypes carrying key loci such as HGGs. This integrated approach will enable a transition from agriculture using chemical inputs to biology-driven farming systems, where crops actively recruit and cooperate with beneficial microbes for nutrition, growth, and protection. In this regard, we already observed that a large number of microbes isolated from natural B. rapa populations can reduce disease on the host plant.
Our next step will be to assess the range of biocontrol of these microbes on other crops important for European farming, such as wheat and maize. We also observed that disease control is higher when ad hoc-designed communities are applied. The ultimate challenge will be to develop broad-spectrum microbial communities or a dedicated process for defining these communities based on the plant genotype targeted, through precision agriculture.
References
Maillet, L. et al. (2025) ‘Plant Genetic Bases Associated With Microbiota Descriptors Shed Light Into a Novel Holobiont Generalist Genes Theory’, Environmental Microbiology, 27(5), e70108. Available at: https://doi.org/10.1111/1462-2920.70108.
PROJECT SUMMARY
The evolutionary processes underlying interactions between hosts and their associated microbes remain a black box that biologists have long sought to illuminate.
A major limitation has been the lack of knowledge about precise molecular events, which are now increasingly accessible through next-generation sequencing technologies. The ERC Starting Grant HOLOE2Plant project leads the way in host-microbiome evolutionary studies, utilising fast-cycling B. rapa plants and synthetic microbial communities. By harnessing high-throughput sequencing, experimental coevolution and advanced modelling, it unravels plant-microbiome interactions under pathogen pressure across multiple plant generations. By validating the holobiont concept and identifying key genetic mechanisms, HOLOE2Plant sets the stage for designing microbial consortia for biocontrol and biostimulation, advancing functional evolutionary science in service of agroecology.
PROJECT LEAD PROFILE
Dr Claudia Bartoli, an evolutionary biologist at INRAE’s LIPME Laboratory in Toulouse, specialises in microbial evolution and plant health. Her research integrates genomic modelling, coevolution experiments and association studies to uncover plant and microbial genes linked to crop health. Driven by agroecology, she aims to develop innovative, microbe-based strategies to prevent crop diseases sustainably.
PROJECT CONTACTS
Claudia Bartoli (Project Coordinator)
Email: claudia.bartoli-kautsky@inrae.fr
FUNDING
This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 101039541.
Funded by the European Union. Views and opinions expressed are, however, those of the author(s) only and do not necessarily reflect those of the European Union or the European Research Council. Neither the European Union nor the granting authority can be held responsible for them.
Figure legends
Figure 1: Plant and microbiota diversity (images are from Maillet et al., 2025). (A) Diversity of natural habitats where B. rapa plants were collected. (B) Principal component analysis on plant SNPs showing the genetic diversity among the populations analysed. (C) Fungal species diversity expressed as Shannon’s index characterised in each population over 2 seasons.
Figure 2: Manhattan plots illustrating Genome–Environment Association (GEA) analysis for relative abundance of the Trichoderma fungi. The red peak of association on chromosome 1 falls in the A01p01520.1_BnaRCC gene, encoding for a cytochrome P450 in Arabidopsis thaliana. The y-axis corresponds to the values of the Lindley process corresponding to the −log10 (p) of the tuning parameter ξ = 3.