Our planet is often called the blue planet, but it could also be called the green planet. That is because photosynthetic life—the primary producers that can convert the energy from the sun into chemical energy—crowds every corner of land where water is available. In the big blue seas, photosynthetic organisms are also widespread and often abundant, though less visible at first sight.
Photosynthetic life has not only visually altered the look of our planet, but it has also allowed life as we know it to evolve. Primary producers are at the base of every food web, sustaining directly or indirectly almost all living organisms. And by releasing oxygen as a by-product of photosynthesis, they have transformed the chemistry of our biosphere. Thanks to an increasingly oxygenated atmosphere and oceans, larger and more complex organisms such as animals could evolve. It is easy to be fooled into thinking that trees and other land plants that we see when we look outside are the main primary producers. In fact, countless forms of algae, often microscopic (e.g. single-cell organisms), collectively produce as much oxygen as all the plants together; in other words, every other breath that you take is the result of algal photosynthesis.
Endosymbiosis: when photosynthesis entered the eukaryotic realm
The broad strokes for how algae have evolved are well established, but the details are far less understood. It all began with what might seem like an improbable event: the merger and subsequent deep cellular integration of photosynthetic bacteria (cyanobacteria, the pioneers of oxygenic photosynthesis) within a single-celled eukaryote. This process, known as primary endosymbiosis, gave rise to the first photosynthetic eukaryotes and thus marked the entry of photosynthesis into the eukaryotic realm. Remarkably, endosymbiosis is not a rare occurrence in nature—think of the symbiosis in coral, among countless other examples—and it stands as one of the most powerful engines of biological innovation.
Most often, endosymbiosis does not lead to the formation of new cellular organelles. However, the close and enduring association between a host cell and a cyanobacterial endosymbiont in primary endosymbiosis gradually transformed the endosymbiont into new bona fide photosynthetic organelles known as plastids (chloroplasts are the best-known example of plastids).
More plastids: a story of endosymbioses
Primary endosymbiosis is an ancient event estimated to have occurred around 1.5 billion years ago or even earlier. The current paradigm states that it was a singular event, in an ancestor of a major group of eukaryotes (a supergroup, akin to a kingdom if you wish) known as Archaeplastida (‘ancient plastids’). Although the general readers may have never heard of Archaeplastida, they will certainly be familiar with some of its members. Indeed, this is where land plants reside in the tree of life, but also the vast diversity of green algae (for example, the lab model Chlamydomonas), red algae (for example, the nori sheet containing your maki), and a more enigmatic, much smaller, algal group called glaucophytes (Figure 1). According to the single primary endosymbiosis hypothesis, all of these archaeplastid groups possess primary plastids of shared origin.
But the fascinating story of plastid evolution does not stop here. First of all, primary plastid endosymbiosis happened again at least once, much more recently, around 100 million years ago in an amoeba called Paulinella (which belongs to another supergroup called Rhizaria). The much younger age of the Paulinella primary plastids means it has retained more clues about how free-living cyanobacteria transformed into organelles, and, as such, these single-celled organisms have become an important model for studying plastid origin.
Moreover, the tree of eukaryotes contains many more algae, often unrelated, that have in fact more complex plastids that are the result of mergers between eukaryotic cells: one alga with primary plastids or already complex plastids, and one heterotrophic cell host. These higher-order plastid endosymbioses are referred to as secondary, tertiary, or even quaternary endosymbioses, and were at the origin of some of the most successful algae that have ever lived on this planet, such as diatoms (members of stramenopiles), dinoflagellates (members of alveolates), or coccolithophorids (members of haptophytes) (Figure 1).
More unknowns than knowns
In PlastidOrigin, a European Research Council-funded project (Horizon 2021 research and innovation programme), we argue that critical data is lacking from the vast ‘hidden’ environmental diversity of microbes to be able to adequately investigate the origin of plastids. Indeed, it is now well appreciated that we have seen, and in the best cases cultured, only a tiny fraction of the immense microbial diversity around us, likely less than 1 %. The rest of the diversity—the great majority—is only known from environmental sequence data (the known unknowns), or often, has not even been sequenced yet (the unknown unknowns). This hidden diversity represents an untapped reservoir of information that can transform our understanding of evolutionary processes, including the endosymbiotic origin of plastids.
One excellent example of how hidden diversity can change how we understand plastid evolution is a phylum named Picozoa, which was a trigger for writing the PlastidOrigin proposal. This group of tiny, elusive microbes has been known for about 2 decades but recently gained attention when my team and colleagues showed it to be a unique archaeplastid lineage (Schön et al. 2021). Using single-cell genomics and phylogenomics, we showed not only that Picozoa is specifically related to red algae (Figure 1), but also confirmed earlier microscopic observations that picozoans lack any trace of a plastid. In other words, picozoans are the first known archaeplastids without a plastid. If you recall what we described previously about Archaeplastida—that they all have primary plastids of shared ancestry, our discovery raises 2 equally interesting possibilities: either Picozoa lost their plastids early in their evolution, which would be unprecedented in a free-living group, or Picozoa never had a plastid. This second possibility would further imply that plastids in eukaryotes were not established by a single primary endosymbiosis, but by multiple ones, and thus that the point of entry of photosynthesis in the eukaryote domain was more complicated than commonly assumed. A textbook changer!
If a group like Picozoa exists, chances are that other equally interesting unknown microbes await discovery. One major aim of PlastidOrigin is precisely that: searching for, and hopefully finding, using a combination of high-throughput environmental sequencing and microscopy, novel, rare, and currently hidden taxa related to known algal groups. And we already found one! This time, we took advantage of the mass of metagenomic data available—when the total DNA of an environment is sequenced—to assemble over 650 new plastid genomes of environmental algae (Jamy et al. 2026). This catalogue of plastid genomes vastly expands the diversity within known major algal groups, often corresponding to specific algae without closely related reference genomes obtained from cultured species. Most interestingly, we also found in this catalogue a novel plastid group, unrelated enough to the known major algal groups that we gave it a new name: leptophytes (Figure 1).
What are the leptophytes? No one has ever seen these algae, or if they have, they did not recognise the importance of what they saw. This is an interesting observation in itself, as algae are among the best-studied microbes. So the fact that there are still unknown major algal groups hiding in plain sight in the sunlit oceans is a testament to how little we know about the diversity of microbes in general. What we can say, based on size fractionation, is that leptophytes are tiny photosynthetic cells, most likely 5 µm or less. We also inferred that this group is globally distributed across all oceans but generally rare, which is probably one of the main reasons leptophytes have escaped visual identification. Excitingly, the phylogenetic position of leptophytes in the eukaryotic tree of life led us to propose new models for the evolution of complex plastids of red algal origin, once again highlighting the importance of discovering new lineages to refine our understanding of biological processes.
The rest of PlastidOrigin
While looking into the unknowns is a big part of PlastidOrigin, this ERC-CoG grant is more than that. Another important aim is the study of Paulinella. As mentioned above, Paulinella experienced the only other known case of primary plastid endosymbiosis. Thus far, almost all research on this group has focused on the few described photosynthetic species, but a larger diversity of closely related uncultured species is known. We often assume that all these species are heterotrophic, but this remains to be confirmed; in fact, we recently described a fourth photosynthetic species (Pardasani et al. 2025) (Figure 2). In PlastidOrigin, we are systematically searching environmental samples to isolate and culture phagotrophic Paulinella species.
This is a risky endeavour, but one that might help to better understand how primary endosymbiosis unfolded in Paulinella by generating high-quality data for close relatives to the plastid-containing species.
Finally, PlastidOrigin will also rely on readily available algal cultures to derive the proteome—the protein content—of plastids and other organelles across Archaeplastida. To do this, we will use a method called hyperplexed localisation of organelle proteins by isotope tagging, or hyperLOPIT, which captures with high spatial resolution the sub-cellular location of proteins. Our main aim is to characterise the evolutionary origin of organellar proteins and the size of the endosymbiotic versus non-endosymbiotic proteomic fractions across Archaeplastida.
Combined, we hope that this project will overcome significant obstacles to understanding how plastids originated, thereby broadly influencing research on endosymbiosis throughout the tree of life. More generally, the methods and approaches we are developing will be applicable to the rest of the microbial eukaryotes. Thus, this project represents a test bed for future global studies aimed at understanding the evolution and ecology of the majority of complex life.
References
Jamy, M. et al. (2026). ‘Identification of a deep-branching lineage of algae using environmental plastid genomes’, Nature Communications, 17, 662. Available at: https://doi.org/10.1038/s41467-025-67401-4.
Pardasani, Y. et al., (2025) ‘Paulinella acadia sp. nov., a New Photosynthetic Species Isolated From a Brackish Beach in British Columbia (Canada)’, Journal of Eukaryotic Microbiology, 72(5), e70040. Available at: https://doi.org/10.1111/jeu.70040.
Schön, M.E. et al., (2021) ‘Single cell genomics reveals plastid-lacking Picozoa are close relatives of red algae’, Nature Communications, 12(1), 6651. Available at: https://doi.org/10.1038/s41467-021-26918-0.
Testing the paradigm of a single plastid origin in eukaryotes (PlastidOrigin)
PROJECT SUMMARY
This multidisciplinary ERC Consolidator project combines single-cell approaches, evolutionary genomics, microscopy, and spatial proteomics to shed new light on the origin and evolution of plastids. We place a strong emphasis on including a better sampled diversity by searching for novel uncultured microbial lineages and using algae in culture, to overcome the current limitation of models of plastid evolution.
PROJECT PARTNERS
PlastidOrigin is based in the Systematic Biology Programme, Department of Organismal Biology, at Uppsala University. Key collaborations include research groups in Spain, France, and Canada.
PROJECT LEAD PROFILE
Fabien Burki was born and raised in Switzerland. After graduating with a PhD from the University of Geneva in 2009, he moved to the University of British Columbia (Canada) for a postdoc. In 2016, Dr Burki established his own research group at Uppsala University (Sweden), where he is now a tenured faculty. Dr Burki has authored over 70 peer-reviewed papers.
PROJECT CONTACTS
Associate Prof. Dr Fabien Burki
Evolutionary Biology Center (EBC) Norbyvägen 18D
752 36 Uppsala, Sweden
Email: fabien.burki@ebc.uu.se
Web: burki-lab.net
FUNDING
This project has received funding from the European Research Council (ERC) under the European Union’s research and innovation programme (Grant agreement No. 101044505).
Funded by the European Union. Views and opinions expressed are, however, those of the author(s) only and do not necessarily reflect those of the European Union or the European Research Council. Neither the European Union nor the granting authority can be held responsible for them.
Figure legends
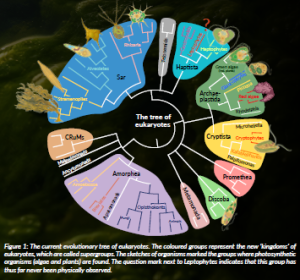
Figure 1: The current evolutionary tree of eukaryotes. The coloured groups represent the new ‘kingdoms’ of eukaryotes, which are called supergroups. The sketches of organisms marked the groups where photosynthetic organisms (algae and plants) are found. The question mark next to Leptophytes indicates that this group has thus far never been physically observed.
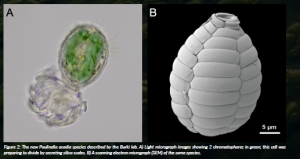
Figure 2: The new Paulinella acadia species described by the Burki lab. A) Light micrograph images showing 2 chromatophores in green; this cell was preparing to divide by secreting silica scales. B) A scanning electron micrograph (SEM) of the same species.